Microwave Imaging for Brain Stroke Detection in Portable Settings
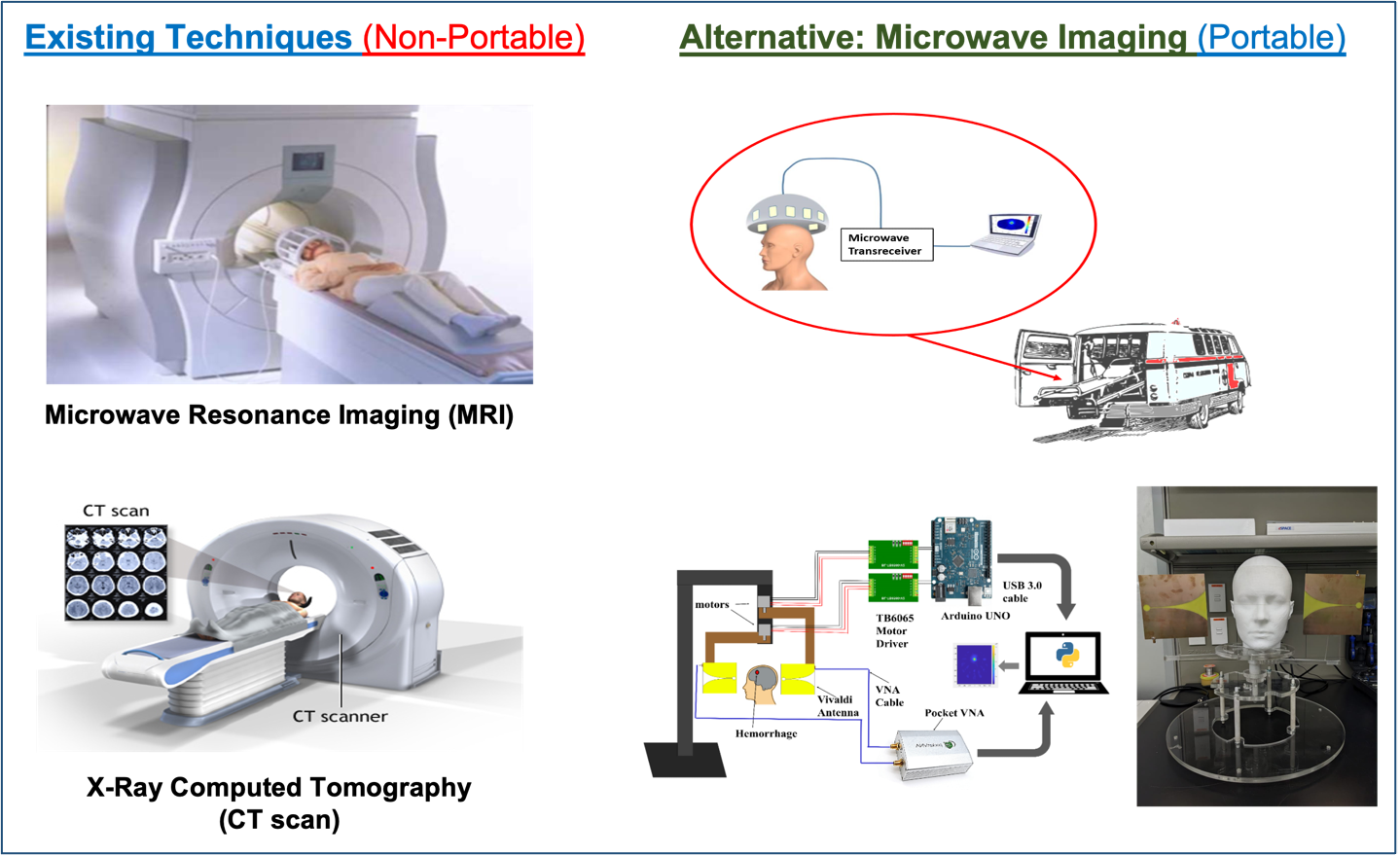
According to the World Health Organization (WHO), stroke is the second leading cause of death and disability. Although there are techniques for the detection and diagnosis of strokes, such as MRI, CT scan, etc., these systems are usually non-portable, available only at tertiary care hospitals, and require that a patient be transported to the imaging device, thus increasing the time taken to apply treatments. ITL has been focusing on developing a Microwave Imaging (MWI) device that would allow local, accurate, rapid, and manageable diagnostic techniques capable of pre-hospital differentiation between different types of stroke. Microwave Imaging (MWI) technology exploits the differences in dielectric properties of healthy tissues and stroke-affected tissues. This technique can reconstruct the head image maps by impinging microwave signals on the head and processing the received scattered signals. The proposed device can be built in compact and portable formats at a relatively low cost than MRI /CT scan and will be easy to use, particularly for non-specialists, can also be used for diagnosis of life-threatening conditions in ambulances or at the scene of injury. This project advances the science and technology in biomedical electronics, microwave imaging, low-cost, non-ionizing, and safe diagnosis. The developed prototype would be of great potential for commercialization in the national health system, health clinics, and hospitals.
AI-enabled Brain–Computer Interface in Biomedical Applications
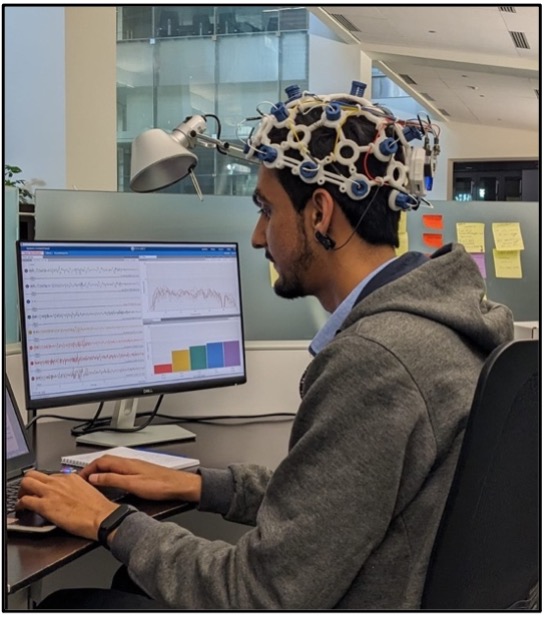

Globally, stroke is recognized as a public health issue and an important challenge for economic development. Stroke is a devastating condition with haemorrhagic or ischemic cause. Ischemic stroke has a worldwide prevalence of 67.6 million (2.7 million annual deaths) and haemorrhagic stroke 15.3 million (2.8 million annual deaths). It is estimated that no more than 1–8% of patients with stroke obtain proper treatment due to pre-hospital delays. At the same time, those who receive the hospital care require continuous monitoring of motor function and brain activity during the rehabilitation process. Therefore, there is imperative to develop technologies that can sustain and work within the complex and fragile environment.
Electrocochleography (ECG) is an important tool for monitoring such patients. Also, stroke severity can be measured for choosing the most optimal rehabilitation program. ITL is exploring the open-source brain-computer interfaces (BCI) for monitoring brain activity. Some special phantoms can be built to mimic various stroke conditions. The data collected from BCI device can then be used to train suitable AI-models for useful decision making in clinical settings.
Automated Blood Cell Counting and Classification using Deep Learning

Medical image processing is becoming one of the most efficient and reliable medical engineering applications due to the major advances in segmentation and computational techniques. Image processing is used to analyze x-ray and blood smear images for early diseases diagnosis such as anemia, kidney tumors, and lung diseases. Clinically, to measure the number of blood cells and evaluate their characteristics, a complete blood count (CBC) is required. In biomedicine, CBC is used to analyze patient’s overall health and detect blood diseases or abnormal immunological reaction by measuring the number and/or morphology of red blood cells (RBCs) and white blood cells (WBCs). Traditional processes for CBC rely on human assistance, where counting and classifying cells are done manually through a microscope. Those techniques are usually inaccurate, time consuming, and may lead to erroneous results as the whole process is subject to human factors. Currently, some hospitals and laboratories start applying sophisticated analyzers which use laser flow cell sensors to detect and count different blood elements and cells automatically. However, those instruments are very expensive and most of laboratories and hospitals in rural places are unable to afford such sophisticated and efficient equipment. Hence, a computer-based solution is required to automate the CBC and reduce its cost.
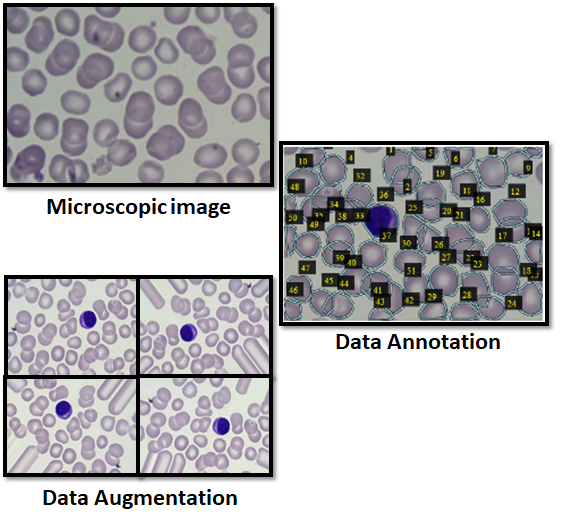

Recently, microscopic image processing has received more attention and has shown impressive improvements in biomedicine to enable diagnostic tests for diverse healthcare applications. Different approaches have been suggested in literature in order to automate RBCs and WBCs counting. Most of those studies focused on images filtering and pre-processing to enhance cells segmentation and counting. Meanwhile, artificial intelligence (AI) is also witnessing an increasing proliferation in biomedicine and researchers are investigating its use in real world and healthcare applications through machine learning techniques and neural network computing. In fact, AI proved its efficiency in bioclinical medicine, molecular medicine, and medical imaging especially in pattern recognition and image segmentation. In this research, a novel framework that automatically counts and classifies different blood cells, i.e., RBCs and WBCs, in a given microscopic blood smear image using a combination of convolutional neural network (CNN), transfer learning, and mask R-CNN techniques. The objective is to apply image segmentation techniques in order to locate, predict boundaries, classify, and count RBCs and WBCs in blood smear images. Unlike previous studies which relied on traditional image processing techniques and image segmentation, in this work, we advocate the use of mask R-CNN for not only its efficiency in object detection and classification but also for its performance in instance segmentation. As a result, it can be utilized to detect different cells blood smears and differentiate between overlapped cells and objects belonging to the same class. In this context, we propose the use of Resnet-101 as a backbone for feature pyramid network model and Microsoft common objects in context (MS-COCO) pre-trained model to initiate the neural network model weights. In addition, data augmentation and regulation techniques are applied to enhance the model detection and reduce the counting error. The obtained results reveal a highly detection rate of different blood cells. In addition, unlike other state-of-the-art techniques, our proposed method has the ability to identify overlapped and faded cells.
