
Abderrazak Chahid
About
Abderrazak Chahid is a PhD candidate in Electrical and Computer Engineering department within CEMSE division. He obtained his Bachelor of Science degree in Electrical Engineering and Power Electronics from Sultan Moulay Slimane University in Morocco. He then got a Master degree in Electrical Engineering from the National School of Applied Sciences in Morocco before obtaining a second Master of Science degree in Embedded Systems from Lorraine University in France.
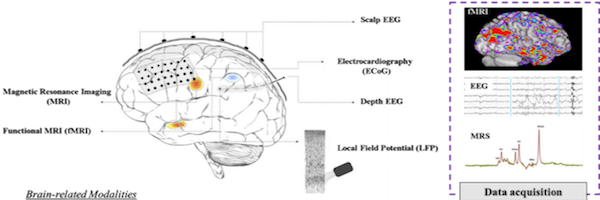
Abderrazak is currently working on developing biomedical signal/image processing: MRS water suppression, MRI image denoising. He is also working on designing signal processing-based feature extraction methods to build machine learning models for biomedical applications: Epileptic spikes detection using MEG/EEG/ECG signals, Hand Gesture detection using sEMG signals.
Research activities
Nowadays, there is a recent call to build a human-independent intelligence which can assist clinicians during medical diagnosis. However, the biomedical records have high dimensionality and are usually contaminated by various types of noises.
In our research, we focus on investigating the use of signal processing tools for biomedical diagnosis.
Biomedical Signal/Image quality enhancement
In order to improve the biomedical signal quality, we designed different algorithms based on a quantum method called Semi-Classical Signal Analysis (SCSA). This method has the ability to decompose the input signal/image into a set of components based on the Eigen-spectrum of the Schrödinger operator.
We targeted different biomedical applications:
- MRI image enhancement using adaptive algorithm based on image segmentation
- MRS residual water suppression using the squared eigenfunctions of the Schrödinger operator
- Complex MRS signal denoising using SCSA-based soft thresholding.
Biomedical Signal/Image classification
In order to assist neurologist and biologist during their daily activity and save the time spent in the manual processing of the records, we need focus on building efficient machine learning models based on the extraction of pertinent and discriminative features for the prediction of biomedical signals. Our models are based on:
- Using Digital Signal Processing (DSP) features
- Improving the raw data quality and reducing feature vector size
- Improving the classifier robustness or sensitivity to noise.
In our different studies, we targeted the following aspects:
- Assist biologists in defining accurately the Poly(A) locations in the DNA sequence
- Assist clinicians in detecting spiky region in the brain using MEG/EEG/ECG signals epileptic diagnosis
- Predicting human cognitive tasks from their corresponding functional Magnetic Resonance Imaging (fMRI) data.
Education Profile
- MSc. Embedded Systems from Lorraine University in France (2016)
- MSc. National School of Applied Sciences in Morocco (2016)
- BSc.Sultan Moulay Slimane University in Morocco (2014)