© 2021 KAUST; Veronica Moraru
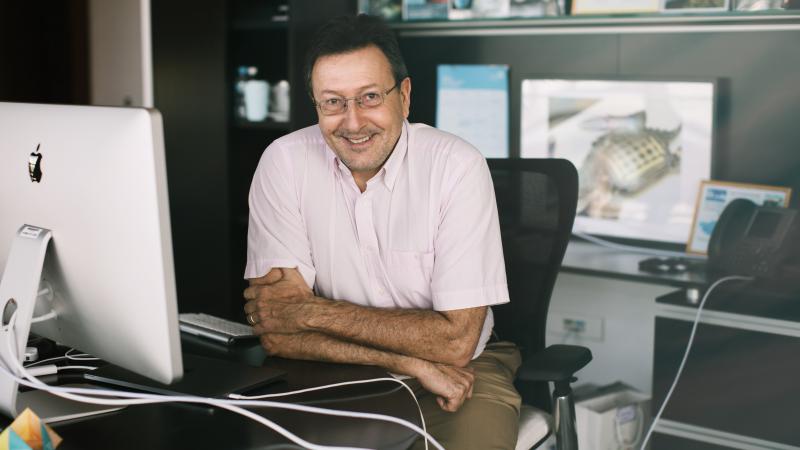
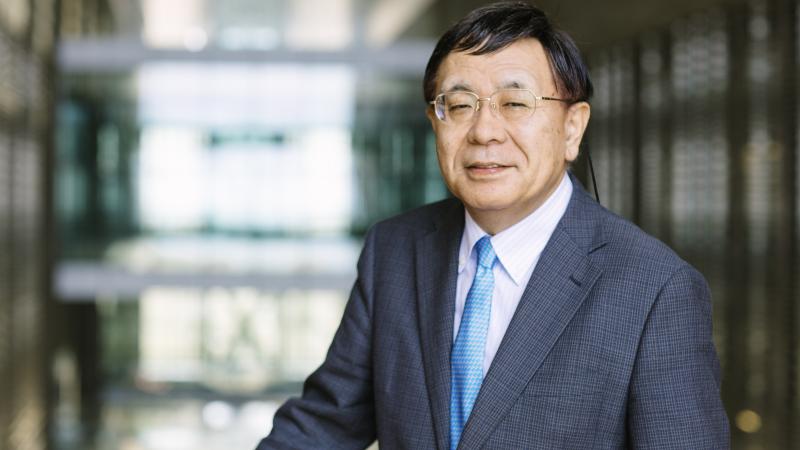
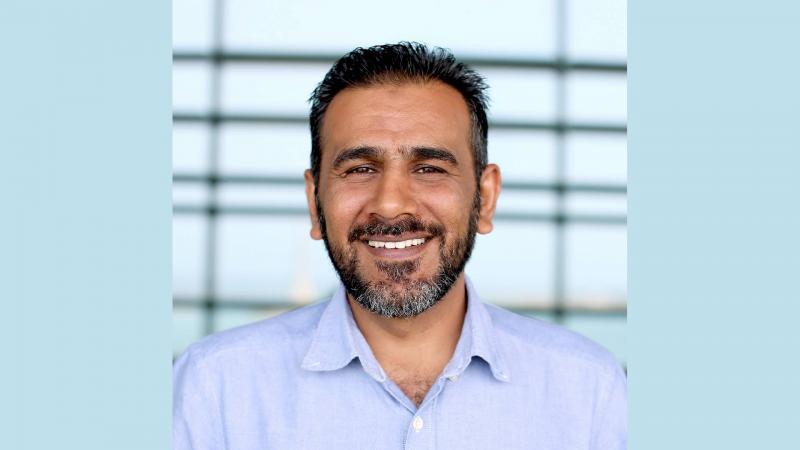
An online platform now gives scientists around the world access to KAUST’s advanced computational resources to improve understanding of the kinds of microbes that exist in different environments, and what they can do. The tool is expected to help researchers identify proteins and enzymes that can be used in agriculture, pharmaceuticals, the energy sector and many other industries.
Preparing bacterial cultures is routine: a scientist takes a sample from a wound, for example, and grows bacteria from it in laboratory petri dishes. The problem is that 99 percent of bacteria in these and other samples cannot be cultured like this in the laboratory. This makes it extremely difficult to discover the estimated one trillion microbes that exist.

To overcome this problem, scientists introduced an approach in 1998 called metagenomics sequencing, where a sample, such as a bucket of seawater, is taken from any environment and then analyzed for DNA. Scientists apply a method called shotgun sequencing that fragments any DNA in the sample into smaller pieces called reads. These metagenomic short reads are then reassembled to identify genes. Over the years, a tremendous amount of microbial sequencing data has been extracted from different environments, but analysis requires state-of-the-art methods, recent reference databases and huge computational prowess.
Read the full article